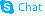Chenodeoxycholic acid
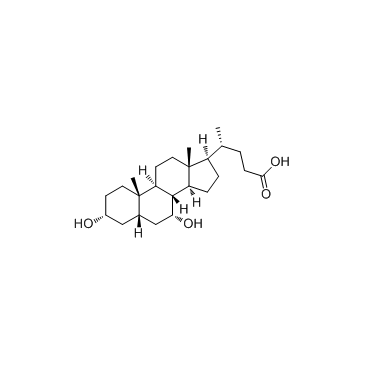
Chenodeoxycholic acid structure
|
Common Name | Chenodeoxycholic acid | ||
|---|---|---|---|---|
| CAS Number | 474-25-9 | Molecular Weight | 392.572 | |
| Density | 1.1±0.1 g/cm3 | Boiling Point | 547.1±25.0 °C at 760 mmHg | |
| Molecular Formula | C24H40O4 | Melting Point | 165-167 °C(lit.) | |
| MSDS | Chinese USA | Flash Point | 298.8±19.7 °C | |
| Symbol |

GHS07 |
Signal Word | Warning | |
Use of Chenodeoxycholic acidChenodeoxycholic Acid is a hydrophobic primary bile acid that activates nuclear receptors (FXR) involved in cholesterol metabolism. |
| Name | chenodeoxycholic acid |
|---|---|
| Synonym | More Synonyms |
| Description | Chenodeoxycholic Acid is a hydrophobic primary bile acid that activates nuclear receptors (FXR) involved in cholesterol metabolism. |
|---|---|
| Related Catalog | |
| Target |
Human Endogenous Metabolite |
| In Vitro | Chenodeoxycholic acid (CDCA) and Deoxycholic acid (DCA) both inhibit 11 beta HSD2 with IC50 values of 22 mM and 38 mM, respectively and causes cortisol-dependent nuclear translocation and increases transcriptionalactivity of mineralocorticoid receptor (MR)[1]. Chenodeoxycholic acid is able to stimulate Ishikawa cell growth by inducing a significant increase in Cyclin D1 protein and mRNA expression through the activation of the membrane G protein-coupled receptor (TGR5)-dependent pathway[2]. Chenodeoxycholic acid (CDCA) induces LDL receptor mRNA levels approximately 4 fold and mRNA levels for HMG-CoA reductase and HMG-CoA synthase two fold in a cultured human hepatoblastoma cell line, Hep G2[3]. Chenodeoxycholic acid-induced Isc is inhibited (≥67%) by Bumetanide, BaCl2, and the cystic fibrosis transmembrane conductance regulator (CFTR) inhibitor CFTRinh-172. Chenodeoxycholic acid-stimulated Isc is decreased 43% by the adenylate cyclase inhibitor MDL12330A and Chenodeoxycholic acid increases intracellular cAMP concentration[4]. Chenodeoxycholic acid treatment activates C/EBPβ, as shown by increases in its phosphorylation, nuclear accumulation, and expression in HepG2 cells. Chenodeoxycholic acid enhances luciferase gene transcription from the construct containing -1.65-kb GSTA2 promoter, which contains C/EBP response element (pGL-1651). Chenodeoxycholic acid treatment activates AMP-activated protein kinase (AMPK), which leads to extracellular signal-regulated kinase 1/2 (ERK1/2) activation, as evidenced by the results of experiments using a dominant-negative mutant of AMPKα and chemical inhibitor[5]. |
| Kinase Assay | Briefly, transfected HEK-293 cells, incubated in charcoal-treated Dulbecco's modified Eagle's medium for 24 h, are washed once with Hanks' solution and resuspended in a buffer containing 100 mM NaCl, 1 mM MgCl2, 1 mM EDTA, 1 mM EGTA, 250 mMsucrose, 20 mM Tris-HCl, pH 7.4. Cells are lysed by freezing in liquid nitrogen. Dehydrogenase activity is measured in a final volume of 20 μL containing the appropriate concentration of bile acid, 30 nCi of [3H]cortisol, and unlabeled cortisol to a final concentrations of 50 nM. The reaction is started by mixing cell lysate with the reaction mixture. Alternatively, endoplasmic reticulum microsomes are prepared from transfected HEK-293 cells and incubated with reaction mixture containing various concentrations of cortisol and CDCA. Incubation proceeded for 20 min, and the conversion of cortisol to cortisone is determined by thin layer chromatography (TLC). Because of the inaccuracy of the TLC method at low conversion rates and the end-product inhibition of 11βHSD2 at conversion rates higher than 60-70%, only conversion rates between 10 and 60% are considered for calculation. The inhibitory constant IC50 is evaluated using the curve-fitting program. Results are expressed as means±S.E. and consist of at least four independent measurements. |
| Cell Assay | The cell viability is analyzed by incubating transfected HEK-293 cells and CHO cells for 1 h with the corresponding concentration of bile acid and staining with trypan blue. The toxicity of bile acids is analyzed using the tetrazolium salt MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) according to the cell proliferation kit I. No significant differences between control and bile acid-treated cells are obtained in both tests. |
| References |
| Density | 1.1±0.1 g/cm3 |
|---|---|
| Boiling Point | 547.1±25.0 °C at 760 mmHg |
| Melting Point | 165-167 °C(lit.) |
| Molecular Formula | C24H40O4 |
| Molecular Weight | 392.572 |
| Flash Point | 298.8±19.7 °C |
| Exact Mass | 392.292664 |
| PSA | 77.76000 |
| LogP | 4.66 |
| Vapour Pressure | 0.0±3.3 mmHg at 25°C |
| Index of Refraction | 1.543 |
| Water Solubility | PRACTICALLY INSOLUBLE |
CHEMICAL IDENTIFICATION
HEALTH HAZARD DATAACUTE TOXICITY DATA
MUTATION DATA
|
| Symbol |

GHS07 |
|---|---|
| Signal Word | Warning |
| Hazard Statements | H315-H319 |
| Precautionary Statements | P305 + P351 + P338 |
| Personal Protective Equipment | Eyeshields;Gloves;type N95 (US);type P1 (EN143) respirator filter |
| Hazard Codes | Xn:Harmful |
| Risk Phrases | R63 |
| Safety Phrases | S22-S24/25-S45-S36/37 |
| RIDADR | NONH for all modes of transport |
| WGK Germany | 2 |
| RTECS | FZ1980000 |
| HS Code | 2918990090 |
| Precursor 9 | |
|---|---|
| DownStream 9 | |
| HS Code | 2918990090 |
|---|---|
| Summary | 2918990090. other carboxylic acids with additional oxygen function and their anhydrides, halides, peroxides and peroxyacids; their halogenated, sulphonated, nitrated or nitrosated derivatives. VAT:17.0%. Tax rebate rate:13.0%. . MFN tariff:6.5%. General tariff:30.0% |
|
Transcriptional Regulation of Cytosolic Sulfotransferase 1C2 by Intermediates of the Cholesterol Biosynthetic Pathway in Primary Cultured Rat Hepatocytes.
J. Pharmacol. Exp. Ther. 355 , 429-41, (2015) Cytosolic sulfotransferase 1C2 (SULT1C2) is expressed in the kidney, stomach, and liver of rats; however, the mechanisms regulating expression of this enzyme are not known. We evaluated transcriptiona... |
|
|
Cheminformatics analysis of assertions mined from literature that describe drug-induced liver injury in different species.
Chem. Res. Toxicol. 23 , 171-83, (2010) Drug-induced liver injury is one of the main causes of drug attrition. The ability to predict the liver effects of drug candidates from their chemical structures is critical to help guide experimental... |
|
|
Translating clinical findings into knowledge in drug safety evaluation--drug induced liver injury prediction system (DILIps).
J. Sci. Ind. Res. 65(10) , 808, (2006) Drug-induced liver injury (DILI) is a significant concern in drug development due to the poor concordance between preclinical and clinical findings of liver toxicity. We hypothesized that the DILI typ... |
| CDCA |
| Chenix |
| MFCD00064142 |
| Chenodex |
| EINECS 207-481-8 |
| Hekbilin |
| Fluibil |
| Ulmenide |
| Chenocol |
| chendol |
| Kebilis |
| anthropododesoxycholicacid |
| Chenodeoxycholic acid |
| 3α,7α-Dihydroxy-5β-cholanic Acid |
| Chendiol |
| 5β-Cholanic Acid-3α,7α-diol |
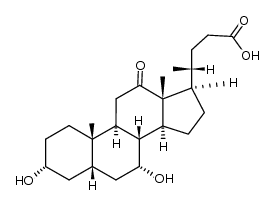 CAS#:2458-08-4
CAS#:2458-08-4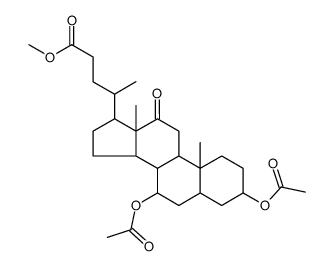 CAS#:28535-81-1
CAS#:28535-81-1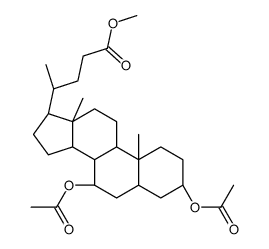 CAS#:2616-71-9
CAS#:2616-71-9 CAS#:2646-38-0
CAS#:2646-38-0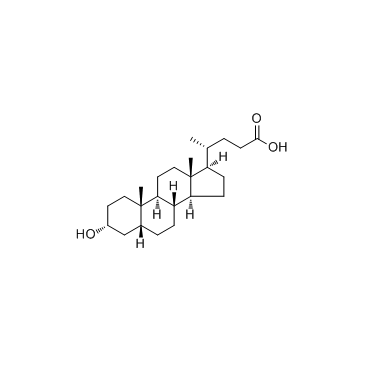 CAS#:434-13-9
CAS#:434-13-9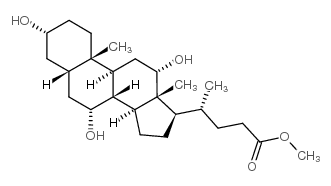 CAS#:1448-36-8
CAS#:1448-36-8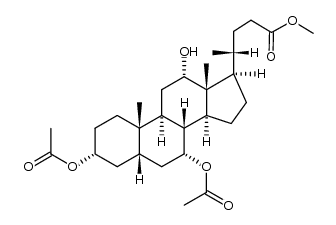 CAS#:3749-87-9
CAS#:3749-87-9 CAS#:71837-87-1
CAS#:71837-87-1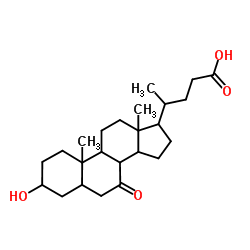 CAS#:4651-67-6
CAS#:4651-67-6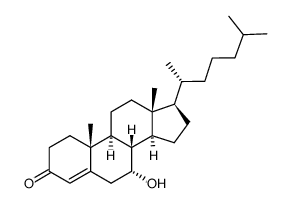 CAS#:3862-25-7
CAS#:3862-25-7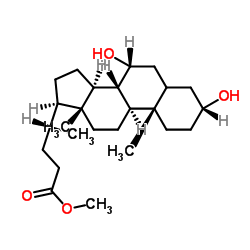 CAS#:3057-04-3
CAS#:3057-04-3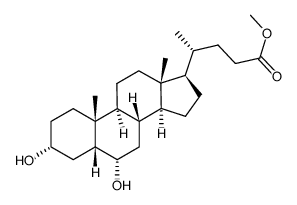 CAS#:2868-48-6
CAS#:2868-48-6 CAS#:10190-22-4
CAS#:10190-22-4 CAS#:2862-62-6
CAS#:2862-62-6 CAS#:6009-98-9
CAS#:6009-98-9 CAS#:16564-43-5
CAS#:16564-43-5