Discovery of candidate disease genes in ENU-induced mouse mutants by large-scale sequencing, including a splice-site mutation in nucleoredoxin.
Melissa K Boles, Bonney M Wilkinson, Laurens G Wilming, Bin Liu, Frank J Probst, Jennifer Harrow, Darren Grafham, Kathryn E Hentges, Lanette P Woodward, Andrea Maxwell, Karen Mitchell, Michael D Risley, Randy Johnson, Karen Hirschi, James R Lupski, Yosuke Funato, Hiroaki Miki, Pablo Marin-Garcia, Lucy Matthews, Alison J Coffey, Anne Parker, Tim J Hubbard, Jane Rogers, Allan Bradley, David J Adams, Monica J Justice
Index: PLoS Genet. 5(12) , e1000759, (2009)
Full Text: HTML
Abstract
An accurate and precisely annotated genome assembly is a fundamental requirement for functional genomic analysis. Here, the complete DNA sequence and gene annotation of mouse Chromosome 11 was used to test the efficacy of large-scale sequencing for mutation identification. We re-sequenced the 14,000 annotated exons and boundaries from over 900 genes in 41 recessive mutant mouse lines that were isolated in an N-ethyl-N-nitrosourea (ENU) mutation screen targeted to mouse Chromosome 11. Fifty-nine sequence variants were identified in 55 genes from 31 mutant lines. 39% of the lesions lie in coding sequences and create primarily missense mutations. The other 61% lie in noncoding regions, many of them in highly conserved sequences. A lesion in the perinatal lethal line l11Jus13 alters a consensus splice site of nucleoredoxin (Nxn), inserting 10 amino acids into the resulting protein. We conclude that point mutations can be accurately and sensitively recovered by large-scale sequencing, and that conserved noncoding regions should be included for disease mutation identification. Only seven of the candidate genes we report have been previously targeted by mutation in mice or rats, showing that despite ongoing efforts to functionally annotate genes in the mammalian genome, an enormous gap remains between phenotype and function. Our data show that the classical positional mapping approach of disease mutation identification can be extended to large target regions using high-throughput sequencing.
Related Compounds
| Structure | Name/CAS No. | Molecular Formula | Articles |
|---|---|---|---|
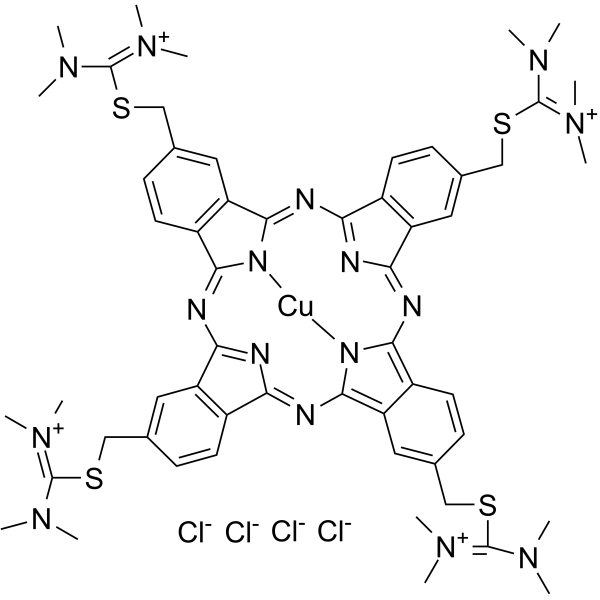 |
C.I. 74240
CAS:33864-99-2 |
C38H30CuN10S+ |
|
Lrp4 domains differentially regulate limb/brain development ...
2015-01-01 [PLoS ONE 10(2) , e0116701, (2015)] |
|
Filamin-interacting proteins, Cfm1 and Cfm2, are essential f...
2014-06-01 [Hum. Mol. Genet. 23(11) , 2953-67, (2014)] |
|
The mych gene is required for neural crest survival during z...
2008-01-01 [PLoS ONE 3(4) , e2029, (2008)] |
|
Genetic analysis of Hedgehog signaling in ventral body wall ...
2011-01-01 [PLoS ONE 6(1) , e16260, (2011)] |
|
Analysis and testing of biological stains--the Biological St...
2002-01-01 [Biotech. Histochem. 77(5&6) , 237-275, (2002)] |