| Structure | Name/CAS No. | Articles |
|---|---|---|
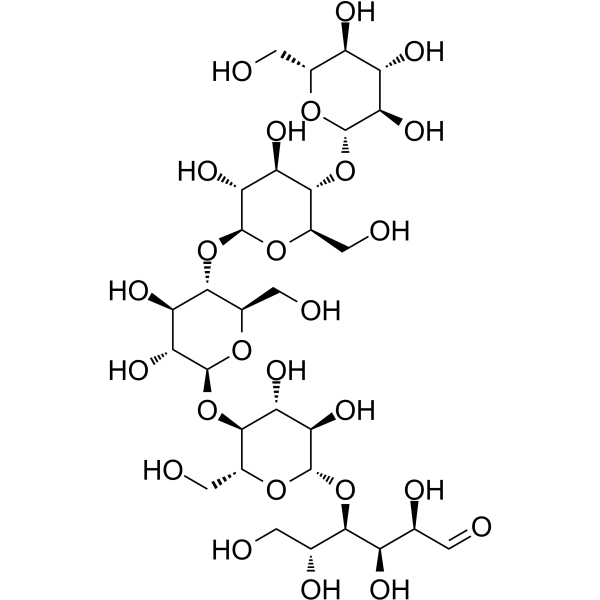 |
Cellopentaose
CAS:2240-27-9 |
|
 |
D-(+)-CELLOTETRAOSE
CAS:38819-01-1 |
|
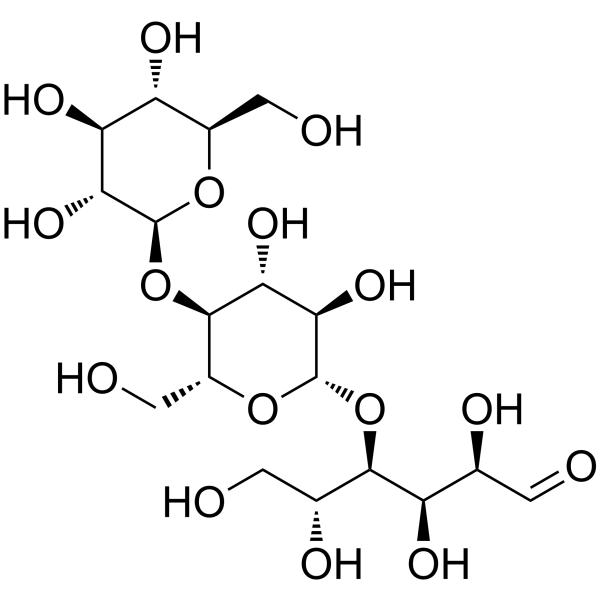 |
D-(+)-CELLOTRIOSE
CAS:33404-34-1 |
|
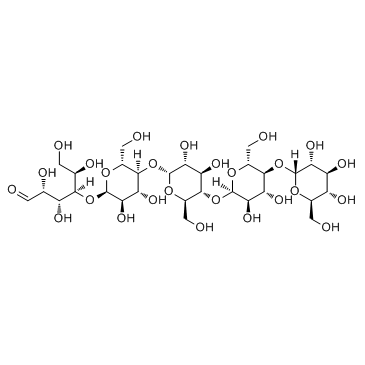 |
Maltopentaose
CAS:34620-76-3 |