Changes of microbial community structures and functional genes during biodegradation of phenolic compounds under high salt condition.
Ping Wang, Yuanyuan Qu, Jiti Zhou
文献索引:J. Environ. Sci. (China) 21(6) , 821-6, (2009)
全文:HTML全文
摘要
The changes of microbial community structures and functional genes during the biodegradation of single phenol and phenol plus p-cresol under high salt condition were explored. It was found that the phenol-fed system (PFS) exhibited stronger degrading abilities and more stable biomass than that of the phenol plus p-cresol-fed system (PCFS). The microbial community structures were revealed by a modern DNA fingerprint technique, ribosomal intergenic spacer analysis (RISA). The results indicated that the microbial community of PFS changed obviously when gradually increased phenol concentration, while PCFS showed a little change. 16S rRNA sequence analysis of the major bands showed that Alcanivorax sp. genus was predominant species during phenolic compounds degradation. Furthermore, amplified functional DNA restriction analysis (AFDRA) on phenol hydroxylase genes showed that the fingerprints were substantially different in the two systems, and the fingerprints were not the same during the different operational periods.
相关化合物
| 结构式 | 名称/CAS号 | 分子式 | 全部文献 |
|---|---|---|---|
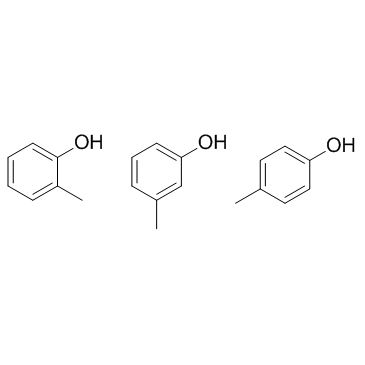 |
混合甲酚
CAS:1319-77-3 |
C7H8O |
|
Purification and characterization of active-site components ...
2008-10-01 [J. Bacteriol. 190(19) , 6493-500, (2008)] |
|
On the decoupling of relaxation modes in a molecular liquid ...
2011-12-08 [J. Phys. Chem. B 115(48) , 13994-9, (2011)] |
|
Cresols utilization by Trametes versicolor and substrate int...
2010-07-01 [Biodegradation 21(4) , 625-35, (2010)] |
|
Co-degradation of phenol and m-cresols by upflow anaerobic s...
2007-01-01 [Water Sci. Technol. 56(7) , 73-9, (2007)] |
|
A simple efficient technique of 'mid-dilution' on-line hemod...
2009-01-01 [Blood Purif. 28(2) , 93-101, (2009)] |