Progress in the bisulfite modification of nucleic acids.
Hikoya Hayatsu, Kazuo Negishi, Yusuke Wataya
Index: Nucleic Acids Symp. Ser. (53) , 217-8, (2009)
Full Text: HTML
Abstract
Bisulfite modification is a principal tool for analyzing DNA methylation, the methyl substitution at position 5 of cytosine residues. Hypermethylation is known to cause silencing of genes, which may result in cell function failures. DNA methylation analysis is therefore a focus of attention in various fields of biological sciences, including even clinical practices for treatment of cancer patients. In 2004, we reported that the bisulfite modification of DNA necessary in this analysis can be speeded up significantly by using a high concentration ammonium bisulfite solution (10 M), in place of traditional sodium bisulfite solution of 5 M concentration. Evaluations on this newer protocol have now come out from several laboratories, showing that this quick process can yield results with greater accuracy compared to those obtainable with widely-practiced low-concentration methods. Another aspect reported here is a study on the desulfonation of uracil-bisulfite adduct to form uracil, the last step of the bisulfite-conversion of cytosine to uracil. Kinetic measurements for the desulfonation of uridine-bisulfite adduct at a near-neutral pH region are described.
Related Compounds
| Structure | Name/CAS No. | Molecular Formula | Articles |
|---|---|---|---|
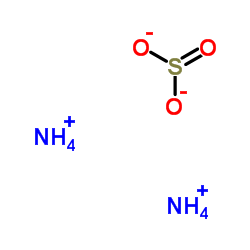 |
ammonium sulfite
CAS:10196-04-0 |
H8N2O3S |
|
13,14-dihydrocoptisine--the genuine alkaloid from Chelidoniu...
2015-03-01 [Phytochemistry 111 , 149-53, (2015)] |
|
Chemistry of bisulfite genomic sequencing; advances and issu...
2007-01-01 [Nucleic Acids Symp. Ser. (51) , 47-8, (2007)] |
|
[Removing the nitrogen from the ammonium sulfite method pape...
2001-07-01 [Huan Jing Ke Xue 22(4) , 117-9, (2001)] |
|
Novel homogeneous catalyst system for the oxidation of conce...
2006-02-28 [J. Hazard. Mater. 129(1-3) , 260-5, (2006)] |
|
Generation, behavior, and toxicity of ammonium sulfite aeros...
1986-01-01 [J. Air Pollut. Control Assoc. 36(1) , 55-9, (1986)] |