Allergenic lipid transfer proteins from plant-derived foods do not immunologically and clinically behave homogeneously: the kiwifruit LTP as a model.
Maria Livia Bernardi, Ivana Giangrieco, Laura Camardella, Rosetta Ferrara, Paola Palazzo, Maria Rosaria Panico, Roberta Crescenzo, Vito Carratore, Danila Zennaro, Marina Liso, Mario Santoro, Sara Zuzzi, Maurizio Tamburrini, Maria Antonietta Ciardiello, Adriano Mari
Index: PLoS ONE 6(11) , e27856, (2011)
Full Text: HTML
Abstract
Food allergy is increasingly common worldwide. Tools for allergy diagnosis measuring IgE improved much since allergenic molecules and microarrays started to be used. IgE response toward allergens belonging to the same group of molecules has not been comprehensively explored using such approach yet.Using the model of lipid transfer proteins (LTPs) from plants as allergens, including two new structures, we sought to define how heterogeneous is the behavior of homologous proteins.Two new allergenic LTPs, Act d 10 and Act c 10, have been identified in green (Actinidia deliciosa) and gold (Actinidia chinensis) kiwifruit (KF), respectively, using clinically characterized allergic patients, and their biochemical features comparatively evaluated by means of amino acid sequence alignments. Along with other five LTPs from peach, mulberry, hazelnut, peanut, mugwort, KF LTPs, preliminary tested positive for IgE, have been immobilized on a microarray, used for IgE testing 1,003 allergic subjects. Comparative analysis has been carried out.Alignment of Act d 10 primary structure with the other allergenic LTPs shows amino acid identities to be in a narrow range between 40 and 55%, with a number of substitutions making the sequences quite different from each other. Although peach LTP dominates the IgE immune response in terms of prevalence, epitope recognition driven by sequence heterogeneity has been recorded to be distributed in a wide range of behaviors. KF LTPs IgE positive results were obtained in a patient subset IgE positive for the peach LTP. Anyhow, the negative results on homologous molecules allowed us to reintroduce KF in patients' diet.The biochemical nature of allergenic molecule belonging to a group of homologous ones should not be taken as proof of immunological recognition as well. The availability of panels of homologous molecules to be tested using microarrays is valuable to address the therapeutic intervention.
Related Compounds
| Structure | Name/CAS No. | Molecular Formula | Articles |
|---|---|---|---|
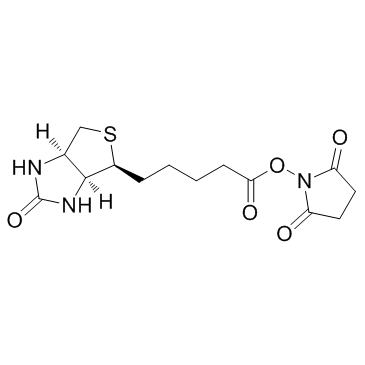 |
Biotin NHS
CAS:35013-72-0 |
C14H19N3O5S |
|
Rapid fluorescent detection of Escherichia coli K88 based on...
2014-07-01 [J. Fluoresc. 24(4) , 1159-68, (2014)] |
|
Xepac protein and IP3/Ca2+ pathway implication during Xenopu...
2015-02-01 [Zygote 23(1) , 99-110, (2015)] |
|
The use of the avidin-biotin complex as a tool in molecular ...
1980-01-01 [Methods Biochem. Anal. New York, NY 26 , 1-45, (1980)] |
|
K. Hoffmann et al.
[J. Am. Chem. Soc. 100 , 3585, (1978)] |
|
On-resin biotinylation of chemically synthesized proteins fo...
1988-05-01 [Anal. Biochem. 170 , 502, (1988)] |