Novel phenylalanine dehydrogenases from Sporosarcina ureae and Bacillus sphaericus. Purification and characterization.
Y Asano, A Nakazawa, K Endo
Index: J. Biol. Chem. 262(21) , 10346-54, (1987)
Full Text: HTML
Abstract
NAD+-dependent phenylalanine dehydrogenases were purified 1,500- and 1,600-fold, and crystallized from Sporosarcina ureae SCRC-R04 and Bacillus sphaericus SCRC-R79a, respectively. The purified enzymes were homogeneous as judged by disc gel electrophoresis. The enzyme from S. ureae has a molecular weight of 305,000, while that of B. sphaericus has a molecular weight of 340,000. Each is probably composed of eight subunits identical in molecular weight. The S. ureae enzyme showed a high substrate specificity in the oxidative deamination reaction acting on L-phenylalanine, while that of B. sphaericus acted on L-phenylalanine and L-tyrosine. The enzymes had lower substrate specificities in the reductive amination reaction acting on alpha-keto acids. The Sporosarcina enzyme acted on phenylpyruvate, alpha-ketocaproate, alpha-keto-gamma-methylthiobutyrate and rho-hydroxyphenylpyruvate. The Bacillus enzyme acted on rho-hydroxyphenylpyruvate, phenylpyruvate, and alpha-keto-gamma-methylthiobutyrate. The enzyme from B. sphaericus catalyzes The enzyme from B. sphaericus catalyzes the transfer of pro-S (B) hydrogen from NADH.
Related Compounds
| Structure | Name/CAS No. | Molecular Formula | Articles |
|---|---|---|---|
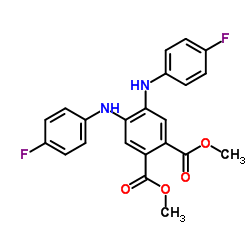 |
l-phenylalanine dehydrogenase
CAS:69403-12-9 |
C22H18F2N2O4 |
|
Fundamental differences in bioaffinity of amino acid dehydro...
2005-03-01 [Anal. Biochem. 338(1) , 102-12, (2005)] |
|
Cloning, sequencing, and expression of Rhodococcus L-phenyla...
1994-06-10 [J. Biol. Chem. 269 , 16203-16211, (1994)] |
|
Phenylalanine dehydrogenase of Bacillus badius. Purification...
1987-10-01 [Eur. J. Biochem. 168(1) , 153-9, (1987)] |
|
Enzymatic phenylalanine estimation for the management of pat...
1991-09-14 [Clin. Chim. Acta 201(1-2) , 95-8, (1991)] |
|
Crystallization of NAD+-dependent phenylalanine dehydrogenas...
1998-03-01 [Acta Crystallogr. D Biol. Crystallogr. 54(Pt 2) , 269-72, (1998)] |