| Structure | Name/CAS No. | Articles |
|---|---|---|
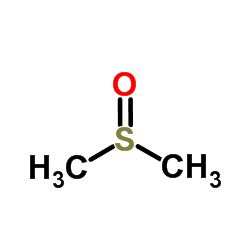 |
Dimethyl sulfoxide
CAS:67-68-5 |
|
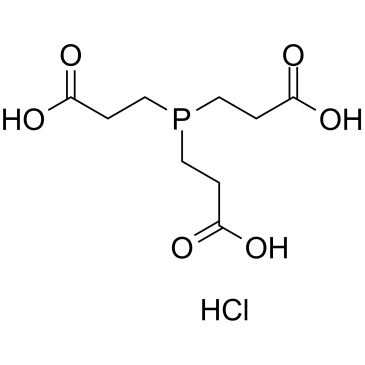 |
Tris(2-carboxyethyl)phosphine Hydrochloride
CAS:51805-45-9 |
|
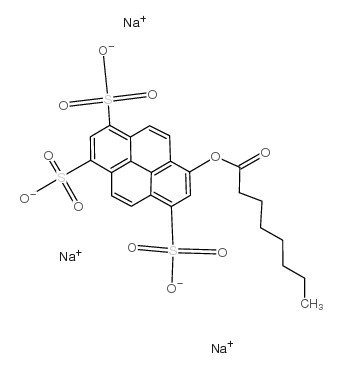 |
8-Octanoyloxypyrene-1,3,6-trisulfonic acid trisodium salt
CAS:115787-84-3 |
|
 |
N,N-Dimethyl-1-naphthylamine
CAS:86-56-6 |